Warning: The `file` argument of `read_csv()` should use `I()` for literal data as of
readr 2.2.0.
# Bad (for example):
read_csv("x,y\n1,2")
# Good:
read_csv(I("x,y\n1,2"))Joins Tutorial
Introduction to joining datasets using dplyr
This document has been adapted and extended by Michael Hallquist and Benjamin Johnson from Jenny Bryan’s dplyr joins tutorial (http://stat545.com/bit001_dplyr-cheatsheet.html). Animations were developed by Garrick Aden-Buie. The goal is to develop an intuition of the four major types of two-table join operations: inner, left, right, and full. We’ll also get into using joins to identify areas of match or mismatch between two datasets (using semi- and anti-joins).
Superheroes table
| name | alignment | gender | publisher |
|---|---|---|---|
| Magneto | bad | male | Marvel |
| Storm | good | female | Marvel |
| Mystique | bad | female | Marvel |
| Batman | good | male | DC |
| Joker | bad | male | DC |
| Catwoman | bad | female | DC |
| Hellboy | good | male | Dark Horse Comics |
Publishers Table
| publisher | yr_founded |
|---|---|
| DC | 1934 |
| Marvel | 1939 |
| Image | 1992 |
Mutating joins: inner, left, right, full
“Mutating” joins combine variables from two datasets on the basis of one or more keys that match between datasets. In the case of these datasets, notice they share the “publisher” column.
N.B.: By default, dplyr will search for common columns across datasets as the matching keys (natural join). If you want to control the process, specify the key using by.
inner join
Require match in both datasets (non-matching rows are dropped)
For those of you who are visual learners, conceptually, imagine the following two simple datasets:
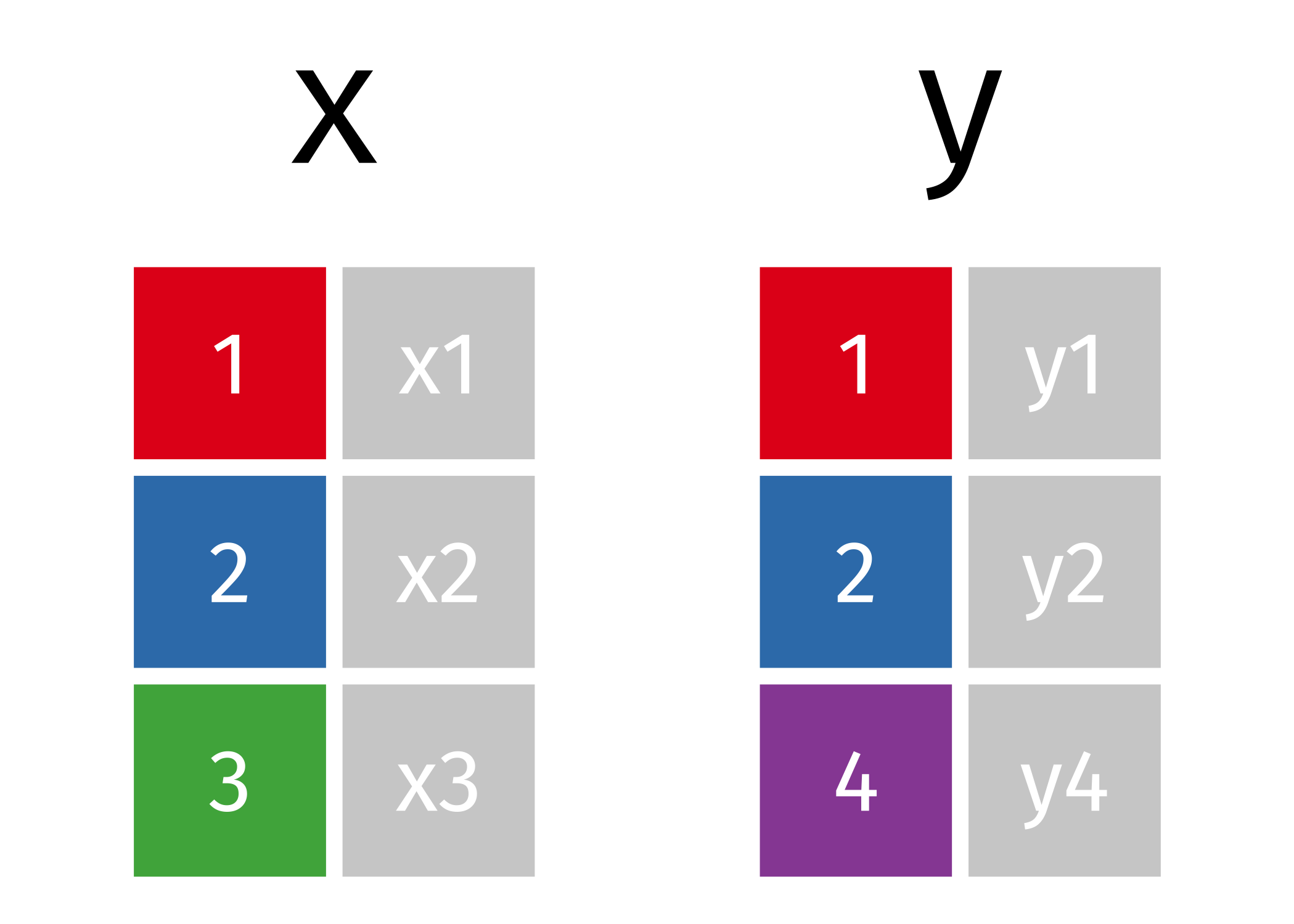
An inner join combines the two datasets and drops the non-matching rows like so:

Let’s try it with our superhero data.
Code
#*NB*: Here, we retain the joined dataset as ijsp
ijsp <- inner_join(x=superheroes, y=publishers)
print(ijsp)# A tibble: 6 × 5
name alignment gender publisher yr_founded
<chr> <chr> <chr> <chr> <dbl>
1 Magneto bad male Marvel 1939
2 Storm good female Marvel 1939
3 Mystique bad female Marvel 1939
4 Batman good male DC 1934
5 Joker bad male DC 1934
6 Catwoman bad female DC 1934Same idea, just explicit declaration of key (i.e., “publisher”)
Code
#note that we've cut the x=, y= as this optional
inner_join(superheroes, publishers, by="publisher")| name | alignment | gender | publisher | yr_founded |
|---|---|---|---|---|
| Magneto | bad | male | Marvel | 1939 |
| Storm | good | female | Marvel | 1939 |
| Mystique | bad | female | Marvel | 1939 |
| Batman | good | male | DC | 1934 |
| Joker | bad | male | DC | 1934 |
| Catwoman | bad | female | DC | 1934 |
Notice both Hellboy (from the superheroes dataset) and Image comics (from the publishers dataset) were dropped.
left join
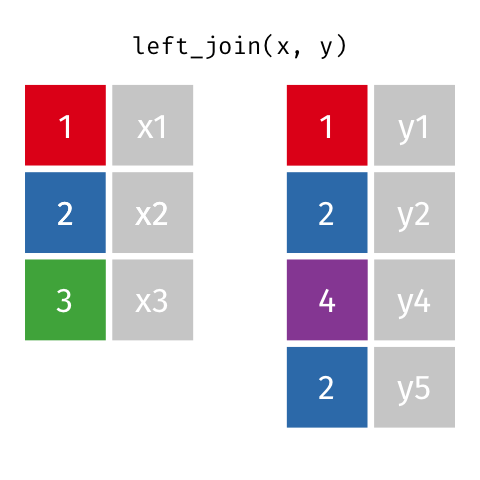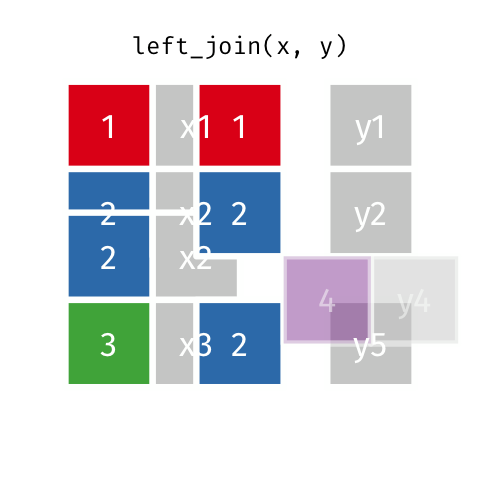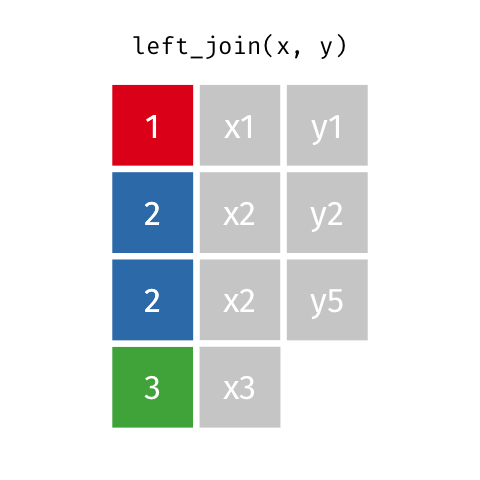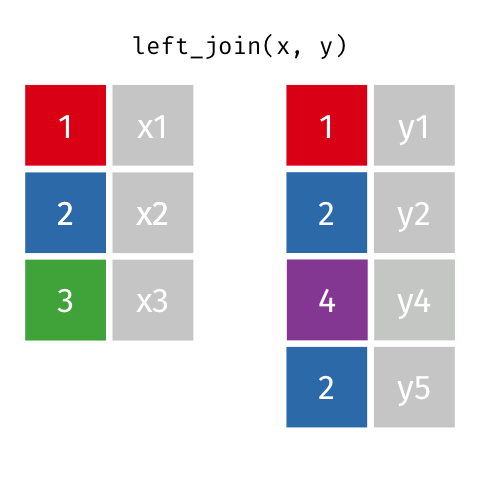
Keep all rows in left-hand ‘x’ dataset (i.e., superheroes). Add columns from publishers where there is a match. Fill in NA for non-matching observations.

Code
left_join(superheroes, publishers, by="publisher")| name | alignment | gender | publisher | yr_founded |
|---|---|---|---|---|
| Magneto | bad | male | Marvel | 1939 |
| Storm | good | female | Marvel | 1939 |
| Mystique | bad | female | Marvel | 1939 |
| Batman | good | male | DC | 1934 |
| Joker | bad | male | DC | 1934 |
| Catwoman | bad | female | DC | 1934 |
| Hellboy | good | male | Dark Horse Comics | NA |
right join
Keep all rows in right-hand ‘y’ dataset (i.e., publishers). Add columns from superheroes where there is a match. Fill in NA for non-matching observations.

Code
# Note the shift to using dplyr piping
# This achieves the same purpose, but may be preferred by those who love pipes
superheroes %>% right_join(publishers, by="publisher")| name | alignment | gender | publisher | yr_founded |
|---|---|---|---|---|
| Magneto | bad | male | Marvel | 1939 |
| Storm | good | female | Marvel | 1939 |
| Mystique | bad | female | Marvel | 1939 |
| Batman | good | male | DC | 1934 |
| Joker | bad | male | DC | 1934 |
| Catwoman | bad | female | DC | 1934 |
| NA | NA | NA | Image | 1992 |
full join
Keep all rows in left-hand ‘x’ (superheroes) and right-hand ‘y’ (publishers) datasets.
Resulting dataset will have all columns of both datasets, but filling in NA for any non-matches on either side (denoted as blank spaces below).

Code
superheroes %>% full_join(publishers, by="publisher")| name | alignment | gender | publisher | yr_founded |
|---|---|---|---|---|
| Magneto | bad | male | Marvel | 1939 |
| Storm | good | female | Marvel | 1939 |
| Mystique | bad | female | Marvel | 1939 |
| Batman | good | male | DC | 1934 |
| Joker | bad | male | DC | 1934 |
| Catwoman | bad | female | DC | 1934 |
| Hellboy | good | male | Dark Horse Comics | NA |
| NA | NA | NA | Image | 1992 |
One-to-many join
Note that when there are non-unique matches, the join adds all possible combinations.
This occurs in a one-to-many join, where a single observation in table A relates to multiple observations in table B. For this scenario, the single table A observation will be replicated onto multiple rows to match the multiple observations in table B.
Let’s say you wanted to examine how mood ratings provided over many days relate to demographic characteristics such as age.
| person | day | mood |
|---|---|---|
| Jeff | 1 | 3 |
| Jeff | 2 | 3 |
| Jeff | 3 | 10 |
| Jeff | 4 | 2 |
| Jeff | 5 | 6 |
| Jeff | 6 | 5 |
| person | bio_sex | age | height_in |
|---|---|---|---|
| Jeff | Male | 15 | 66 |
| Ping | Male | 18 | 71 |
| Karla | Female | 19 | 60 |
| person | bio_sex | age | height_in | day | mood |
|---|---|---|---|---|---|
| Jeff | Male | 15 | 66 | 1 | 3 |
| Jeff | Male | 15 | 66 | 2 | 3 |
| Jeff | Male | 15 | 66 | 3 | 10 |
| Jeff | Male | 15 | 66 | 4 | 2 |
| Jeff | Male | 15 | 66 | 5 | 6 |
| Jeff | Male | 15 | 66 | 6 | 5 |
| Ping | Male | 18 | 71 | 1 | 4 |
| Ping | Male | 18 | 71 | 2 | 6 |
| Ping | Male | 18 | 71 | 3 | 9 |
| Ping | Male | 18 | 71 | 4 | 10 |
| Ping | Male | 18 | 71 | 5 | 5 |
| Ping | Male | 18 | 71 | 6 | 3 |
| Karla | Female | 19 | 60 | 1 | 9 |
| Karla | Female | 19 | 60 | 2 | 9 |
| Karla | Female | 19 | 60 | 3 | 9 |
| Karla | Female | 19 | 60 | 4 | 3 |
| Karla | Female | 19 | 60 | 5 | 8 |
| Karla | Female | 19 | 60 | 6 | 10 |
Notice how the demographic variables are replicated on rows for each day of mood ratings. If both tables have duplicates for the key, joins can produce many-to-many combinations and a large increase in rows.
Filtering joins: semi_join and anti_join
Filtering joins use specific criteria to identify observations (rows) from one table that exist or don’t exist in another table.
These joins are typically used for diagnosing mismatch between two overlapping datasets. That is, filtering joins are most useful for data quality assurance checks.
semi_join
retain observations (rows) in x that match in y

Observations in superheroes that match in publishers
Notice that this is different from the left_join shown above as the data from y is not kept. That is the fundamental difference between ‘mutating joins’ (e.g., left_join) and ‘filtering joins’ (e.g., semi_join).
Code
semi_join(superheroes, publishers, by="publisher")| name | alignment | gender | publisher |
|---|---|---|---|
| Magneto | bad | male | Marvel |
| Storm | good | female | Marvel |
| Mystique | bad | female | Marvel |
| Batman | good | male | DC |
| Joker | bad | male | DC |
| Catwoman | bad | female | DC |
Observations in publishers that match in superheroes
Code
semi_join(publishers, superheroes, by="publisher")| publisher | yr_founded |
|---|---|
| DC | 1934 |
| Marvel | 1939 |
This can be useful if you have a dataset of your data of interest and another dataset that indicates which of your participants/observations you want to remove or filter out.
anti_join
observations in x that are not matched in y Note that this is similar to setdiff in base R

Observations in superheroes that don’t match in publishers
Code
anti_join(superheroes, publishers, by="publisher")| name | alignment | gender | publisher |
|---|---|---|---|
| Hellboy | good | male | Dark Horse Comics |
Observations in publishers that don’t match in superheroes
Code
publishers %>% anti_join(superheroes, by="publisher")| publisher | yr_founded |
|---|---|
| Image | 1992 |
This can be useful if you are trying to identify extra participants/observations that may have snuck into one dataset (x) or been deleted in another (y).
Joining multiple datasets
Joining can be done repeatedly across multiple datasets. The following code, for instance, joins datasets two at a time from left to right in the list. The result of a two-table join becomes the ‘x’ dataset for the next join of a new dataset ‘y’.
Code
set.seed(123)
df1 <- data.frame(id=1:10, x=rnorm(10), y=runif(10))
df2 <- data.frame(id=1:11, z=rnorm(11), a=runif(11))
df3 <- data.frame(id=2:10, b=rnorm(9), c=runif(9))
print(df1) id x y
1 1 -0.56047565 0.8895393
2 2 -0.23017749 0.6928034
3 3 1.55870831 0.6405068
4 4 0.07050839 0.9942698
5 5 0.12928774 0.6557058
6 6 1.71506499 0.7085305
7 7 0.46091621 0.5440660
8 8 -1.26506123 0.5941420
9 9 -0.68685285 0.2891597
10 10 -0.44566197 0.1471136Code
print(df2) id z a
1 1 1.7869131 0.79892485
2 2 0.4978505 0.12189926
3 3 -1.9666172 0.56094798
4 4 0.7013559 0.20653139
5 5 -0.4727914 0.12753165
6 6 -1.0678237 0.75330786
7 7 -0.2179749 0.89504536
8 8 -1.0260044 0.37446278
9 9 -0.7288912 0.66511519
10 10 -0.6250393 0.09484066
11 11 -1.6866933 0.38396964Code
print(df3) id b c
1 2 -0.5996083 0.6680556
2 3 -0.1294107 0.4176468
3 4 0.8867361 0.7881958
4 5 -0.1513960 0.1028646
5 6 0.3297912 0.4348927
6 7 -3.2273228 0.9849570
7 8 -0.7717918 0.8930511
8 9 0.2865486 0.8864691
9 10 -1.2205120 0.1750527Code
dftemp <- full_join(df1, df2, by="id")
dffinal <- full_join(dftemp, df3, by="id")
#alternative way to combine that avoids temporary variables
Reduce(function(x, y) full_join(x, y, by="id"), list(df1, df2, df3))| id | x | y | z | a | b | c |
|---|---|---|---|---|---|---|
| 1 | -0.5604756 | 0.8895393 | 1.7869131 | 0.7989248 | NA | NA |
| 2 | -0.2301775 | 0.6928034 | 0.4978505 | 0.1218993 | -0.5996083 | 0.6680556 |
| 3 | 1.5587083 | 0.6405068 | -1.9666172 | 0.5609480 | -0.1294107 | 0.4176468 |
| 4 | 0.0705084 | 0.9942698 | 0.7013559 | 0.2065314 | 0.8867361 | 0.7881958 |
| 5 | 0.1292877 | 0.6557058 | -0.4727914 | 0.1275317 | -0.1513960 | 0.1028646 |
| 6 | 1.7150650 | 0.7085305 | -1.0678237 | 0.7533079 | 0.3297912 | 0.4348927 |
| 7 | 0.4609162 | 0.5440660 | -0.2179749 | 0.8950454 | -3.2273228 | 0.9849570 |
| 8 | -1.2650612 | 0.5941420 | -1.0260044 | 0.3744628 | -0.7717918 | 0.8930511 |
| 9 | -0.6868529 | 0.2891597 | -0.7288912 | 0.6651152 | 0.2865486 | 0.8864691 |
| 10 | -0.4456620 | 0.1471136 | -0.6250393 | 0.0948407 | -1.2205120 | 0.1750527 |
| 11 | NA | NA | -1.6866933 | 0.3839696 | NA | NA |
Alternative using pipeline (less extensible)
Code
mergedf <- df1 %>% full_join(df2, by="id") %>% full_join(df3, by="id")