set.seed(859)
n_points <- 160
true_angle <- 34 * pi / 180
rotation_mat <- matrix(
c(cos(true_angle), -sin(true_angle),
sin(true_angle), cos(true_angle)),
nrow = 2,
byrow = TRUE
)
raw_cloud <- matrix(rnorm(n_points * 2), ncol = 2) %*%
diag(c(2.8, 0.9)) %*%
rotation_mat
toy_df <- as_tibble(scale(raw_cloud, center = TRUE, scale = FALSE))
names(toy_df) <- c("x1", "x2")
toy_pca <- prcomp(toy_df, center = FALSE, scale. = FALSE)
pc_axes <- tibble(
axis = c("PC1", "PC1", "PC2", "PC2"),
x = c(-1, 1, -1, 1) * c(rep(sqrt(toy_pca$sdev[1]^2) * 3.4, 2), rep(sqrt(toy_pca$sdev[2]^2) * 3.4, 2)) *
c(toy_pca$rotation[1, 1], toy_pca$rotation[1, 1], toy_pca$rotation[1, 2], toy_pca$rotation[1, 2]),
y = c(-1, 1, -1, 1) * c(rep(sqrt(toy_pca$sdev[1]^2) * 3.4, 2), rep(sqrt(toy_pca$sdev[2]^2) * 3.4, 2)) *
c(toy_pca$rotation[2, 1], toy_pca$rotation[2, 1], toy_pca$rotation[2, 2], toy_pca$rotation[2, 2])
) %>%
mutate(endpoint = rep(c("start", "end"), 2))
ggplot(toy_df, aes(x = x1, y = x2)) +
geom_hline(yintercept = 0, linewidth = 0.3, color = "grey80") +
geom_vline(xintercept = 0, linewidth = 0.3, color = "grey80") +
geom_point(color = "#1f78b4", alpha = 0.75, size = 2.2) +
geom_segment(
data = pc_axes %>% filter(axis == "PC1") %>% tidyr::pivot_wider(names_from = endpoint, values_from = c(x, y)),
aes(x = x_start, y = y_start, xend = x_end, yend = y_end),
inherit.aes = FALSE,
color = "#d95f02",
linewidth = 1.2
) +
geom_segment(
data = pc_axes %>% filter(axis == "PC2") %>% tidyr::pivot_wider(names_from = endpoint, values_from = c(x, y)),
aes(x = x_start, y = y_start, xend = x_end, yend = y_end),
inherit.aes = FALSE,
color = "#7570b3",
linewidth = 1.1
) +
annotate("label", x = pc_axes$x[2], y = pc_axes$y[2], label = "PC1", fill = "#d95f02", color = "white") +
annotate("label", x = pc_axes$x[4], y = pc_axes$y[4], label = "PC2", fill = "#7570b3", color = "white") +
labs(
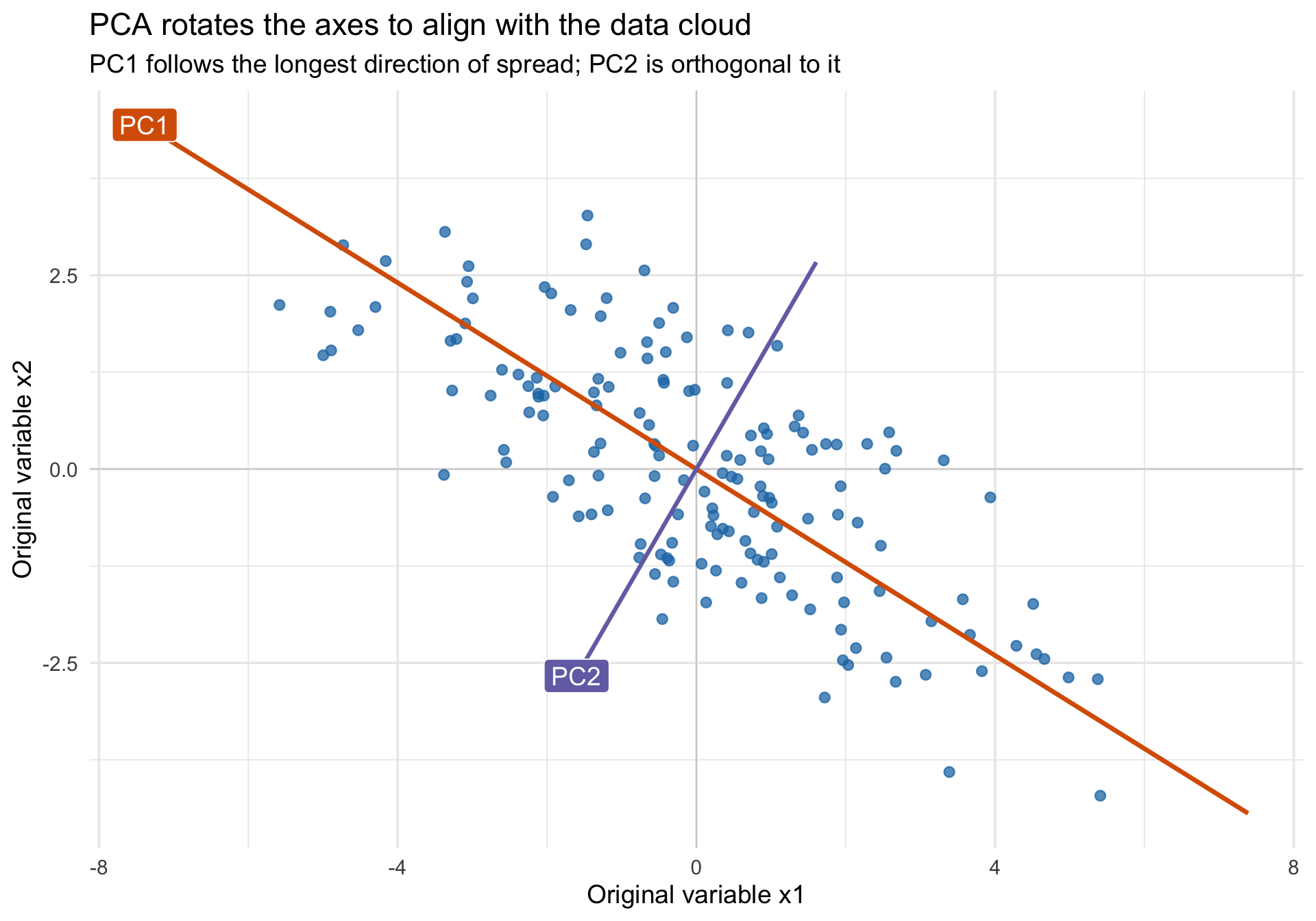
title = "PCA rotates the axes to align with the data cloud",
subtitle = "PC1 follows the longest direction of spread; PC2 is orthogonal to it",
x = "Original variable x1",
y = "Original variable x2"
) +
theme_minimal(base_size = 14)